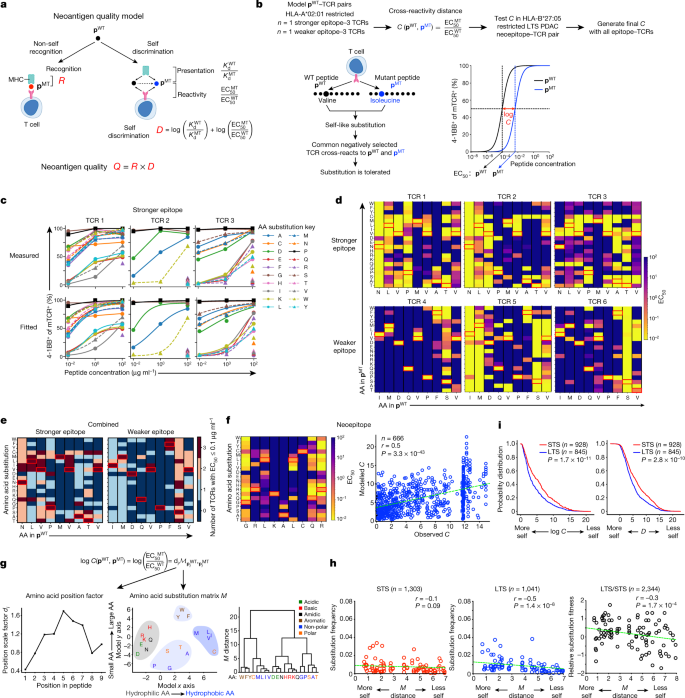
a, Neoantigen high quality mannequin. b, The mannequin and experimental method to estimate cross-reactivity distance C. c, d, Measured (high) and fitted (backside) pMT–TCR activation curves (c, amino acid (AA) place 4), and activation warmth maps (d, all amino acid positions) for stronger and weaker pWT–TCR pairs. e, Composite pMT–TCR EC50 values of all stronger and weaker pWT–TCR pairs. f, pMT–TCR activation warmth map and noticed versus modelled C(pWT, pMT) for the HLA-B*27:05-restricted pWT–TCR pair. n signifies the variety of single-amino-acid-substituted pWT, pMT and pMT, pMT pairs. g, Cross-reactivity distance mannequin C and dendrogram of agglomerative clustering of substitution matrix M. h, Noticed amino acid substitution frequency versus matrix M-defined substitution distance in major and recurrent STS and LTS PDACs. M distance is the matrix M-defined amino acid distance from g. Circles point out substituted residues. n signifies the variety of substitutions. i, Cumulative likelihood distributions of log(C) and D. n signifies the variety of neoantigens. The crimson rectangles within the warmth maps point out amino acids in pWT. The inexperienced line is a linear regression match. Warmth maps are ordered in accordance with the amino acid order within the dendogram in g. P values had been decided utilizing two-tailed Pearson correlation (f and h) and two-sided Kolmogorov–Smirnov assessments (i).
Comments
0 comments